Genetic recombination : understanding the mechanisms / Harold L.K. Whitehouse.
- Harold Leslie Keer Whitehouse
- Date:
- [1982]
Licence: Attribution-NonCommercial-NoDerivatives 4.0 International (CC BY-NC-ND 4.0)
Credit: Genetic recombination : understanding the mechanisms / Harold L.K. Whitehouse. Source: Wellcome Collection.
302/448 page 284
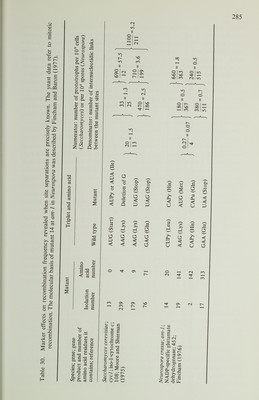
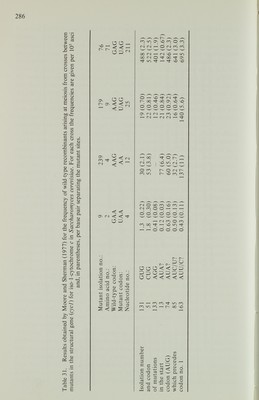
![favour of wild type was found by McKnight, Cardillo and Sherman (1981) with a deletion mutant, H3, of cyc7, the structural gene for iso-2-cytochrome c in S. cerevisiae. In this mutant about 5000 nucleotide pairs are deleted, encompassing part of the regulatory region of cyc7 and two adjacent genes. In 294 tetrads, there were 10 with conversion of H3 to wild type and none with conversion to mutant. Seven of the conversion tetrads were from a cross of H3 with an allele, H2. The H2 mutation was caused by the insertion of about 5500 nucleotide pairs of DNA into the regulatory region of cyc7 not more than 150 nucleotides from the site of H3. In all the seven tetrads the deletion and insertion mutants showed co conversion. Surprisingly, the insertion did not show disparity in the direction of conversion despite its similar size and proximity to H3. From crosses of H2 with wild type or with H3 (omitting the co-conversion asci), there were six tetrads with conversion of H2 to wild type and four with conversion to mutant. More precise information about mismatch correction and other marker effects can be obtained if the gene product has been sequenced in wild-type and mutant strains. The amino acid sequence of iso-1-cytochrome c of S. cerevisiae has been determined and the molecular basis of a number of mutants in cycl , its structural gene, has been established. Moore and Sherman (1975) investigated the recom bination frequencies of eve] mutants. Some of their results are given in Table 30. It is evident that mutants 13 and 239 show an extraordinarily high frequency of mitotic recombination compared with other combinations of alleles. A similar result was obtained with X-ray- and ultraviolet-induced mitotic recombination, and to a lesser extent with meiotic recombination. Subsequently, the investigation was extended to include other mutants mapping in the same part of the gene (Moore and Sherman, 1977). The results were remarkable (Table 31). The dele tion frameshift mutant 239 gave a relatively high frequency of recombination compared with transversions 179 and 76. This result would be accounted for if 239 behaved like deletion frameshifts in Ascobolus and Sordaria and showed con version to wild type more often than to mutant. The probable transversion 163 also gave a relatively high frequency of recombination with alleles, in several instances much higher than was found with other base substitutions at the same site (mutants 13, 74 and 85). Moore and Sherman (1977) concluded that the nature of the mismatched bases influences the recombination process, but not in a way that can be simply interpreted. It seems as if the two mismatches may interact in complex ways to influence the outcome of the recombination event. Thus, the high frequency of mitotic recombination shown by mutants 13 and 239 cannot be predicted from the results of crosses of either of these mutants with other alleles (Table 30). Marker effects apparently related to mismatch correction have also been demonstrated in Schizosaccharomyces pombe. Hofer et al. (1979) found that two opal suppressors, in the sup3 and sup9 genes, respectively, each arose by mutation at the anticodon site of a gene for a transfer RNA for serine. Each mutation was from TGA to TCA and both enhanced the local frequency of meiotic recombina tion five- to 10-fold. Tetrad analysis by Thuriaux et al. (1980) revealed that both mutants gave a high frequency of postmeiotic segregation in one-factor crosses](https://iiif.wellcomecollection.org/image/b18020768_0303.JP2/full/800%2C/0/default.jpg)